Methods
GAP is an integrative, general-purpose framework for deriving a quantitative measure of gene similarity, which is relevant to a wide range of bioinformatics applications from gene clustering and phenotype and protein interactions predictions to interaction network modeling, and pharmacology analysis.
In the current implementation, GAP combines pathway information from many different public online databases, Gene Ontology annotations as well as drug and disease associations from scientific literature.
GAP extends previous computational methods by including a semantic gene similarity measure which exploits the available functional gene annotations. More specifically, the annotations considered in this implementation of GAP come from the Gene Ontology and the strategy that takes advantage of them for assessing the similarity between genes is based on information theoretic principles and exploits the knowledge encoded in the ontology.
GAP, therefore, benefits from a semantic mechanism of assigning gene similarity which processes implicit shared association evidences to identify and prioritize novel predictions of relationships between genes and between their products.
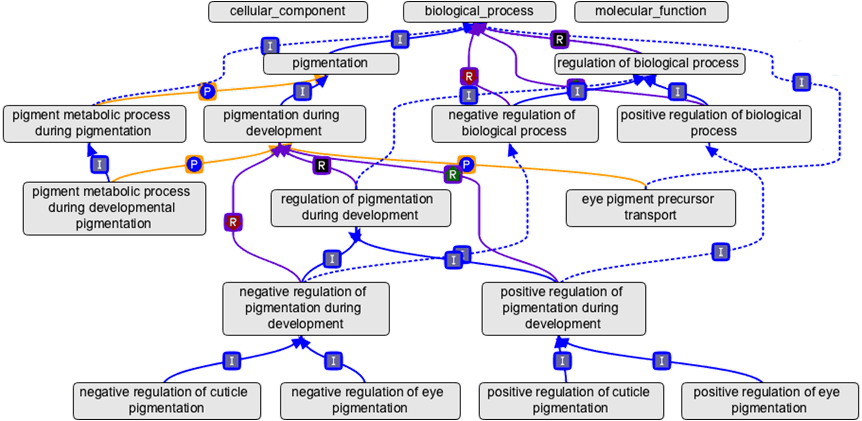
SOURCE: Gene Ontology Annotation Database